|
Clinician's corner |
|
Back to main page |
|
Programmer's corner |
| So, you use WinBUGS a lot? Want more? |
|
Patrick Blisle Division of Clinical Epidemiology McGill University Health Center Montreal, Quebec CANADA patrick.belisle@rimuhc.ca Last modification: 14 jun 2017
|
|
|
Version 1.0.2 (December 2012)
gamma.parms.from.quantiles
[R] Computing Gamma distribution parameters
| [ | gamma.parms.from.quantiles is an R function developped to compute Gamma distribution parameters fitting specified quantiles and cumulative probabilities. | ] |
Menu
Top
Syntax
gamma.parms.from.quantiles <- function(q, p=c(0.025,0.975), precision=0.001, derivative.epsilon=1e-3, start.with.normal.approx=T, start=c(1.1,1.1), plot=F, plot.xlim=numeric(0))
Top
gamma.parms.from.quantiles arguments
| Argument | Value | Comment |
| q | Vector of quantiles; | Vector of length 2; |
| p | Vector of probabilities; |
Vector of length 2. Default is (0.025, 0.975). |
| precision | Desired precision in Gamma distribution parameters; | The algoritm in gamma.parms.from.quantiles is iterative: it will stop when an iteration leads to (shape, rate) estimates that do not differ from estimates obtained in last iteration by more than precision. |
| derivative.epsilon | Small value used for ε in the approximation of the derivatives of a few functions in the algorithm; | A few functions (f) have derivates (f'(x)) that are approximated by (f(x+ε)) - f(x))/ε) in the algorithm, where ε = derivative.epsilon. If the algorithm fails to return correct Gamma distribution parameters, you may want to try again with smaller values than the default (1e-3) for this parameter. |
| start.with.normal.approx | Logical. Whether or not to start the iterative process with values obtained from a normal approximation to the Gamma distribution; | Default is T. |
| start | Vector (of length 2) of initial values for Gamma distribution parameters. | Ignored when start.with.normal.approx is True. |
| plot | Logical. Whether or not to plot the resulting Gamma distribution, along with tail probabilities obtained. | Default is F. |
| plot.xlim |
Lower and upper limits of the plotted area.
◊ Numeric vector of length 2. |
Default value is to include points well beyond the target quantiles specified in q and should fit most needs.
Irrelevant when plot = F. |
Top
Output
| Output dimension | Value | Comment |
| $shape $rate $scale |
parameters of the Gamma distribution; | $scale = 1 / $rate |
| $last.change | last change in $scale and $rate in the iterative algorithm; | |
| $niter | number of iterations run before convergence; | |
| $q | quantiles; | |
| $p | probabilities; | |
| $p.check | a quick validation: values for $scale and $rate returned lead to cumulative probabilities $p.check at the quantiles $q. |
Top
Example
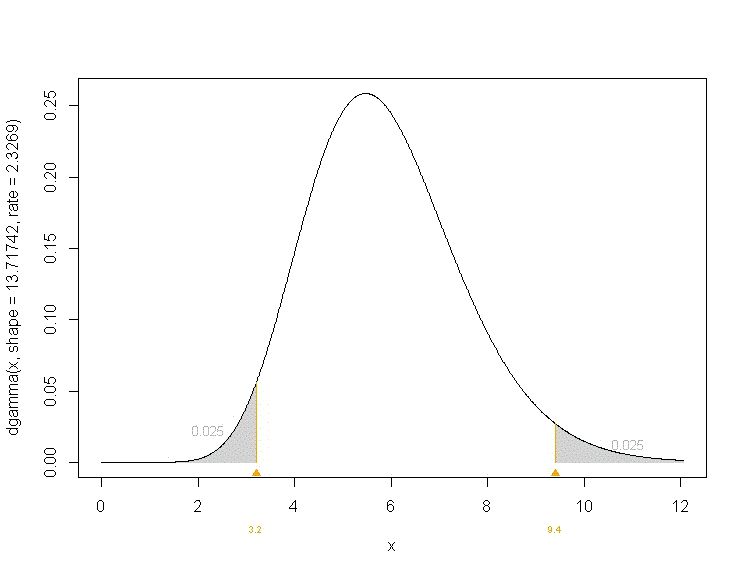
The code below reproduces the input and output of an R session involving a call to gamma.parms.from.quantiles. The use of option plot=T produces the plot attached below (default is to not produce the plot).
The output shows that a Gamma distribution with shape parameter shape=13.72 and rate=2.33 lies in range (3.2, 9.4) with probability 95%, with equal probability tails.
> source('c:/path/GammaParmsFromQuantiles.R') # load file where gamma.parms.from.quantiles R function is defined
> gamma.parms.from.quantiles(c(3.2,9.4), plot=T)
> gamma.parms.from.quantiles(c(3.2,9.4), plot=T)
|
$shape [1] 13.71742 $rate [1] 2.326902 $scale [1] 0.429756 $last.change [1] -3.909504e-04 -6.276569e-05 $niter [1] 5 $q [1] 3.2 9.4 $p [1] 0.025 0.975 $p.check [1] 0.02500001 0.97499999 |
|
Use in a Bayesian context
Gamma densities are frequently used as prior distributions in Bayesian analysis, as they are the conjugate density for precision (the inverse of the variance) in normal likelihoods.
Top
Code
gamma.parms.from.quantiles <- function(q, p=c(0.025,0.975),
precision=0.001, derivative.epsilon=1e-3, start.with.normal.approx=T, start=c(1.1, 1.1), plot=F, plot.xlim=numeric(0))
{
# Version 1.0.2 (December 2012)
#
# Function developed by
# Lawrence Joseph and Patrick B�lisle
# Division of Clinical Epidemiology
# Montreal General Hospital
# Montreal, Qc, Can
#
# patrick.belisle@rimuhc.ca
# http://www.medicine.mcgill.ca/epidemiology/Joseph/PBelisle/GammaParmsFromQuantiles.html
#
# Please refer to our webpage for details on each argument.
f <- function(x, theta){dgamma(x, shape=theta[1], rate=theta[2])}
F.inv <- function(x, theta){qgamma(x, shape=theta[1], rate=theta[2])}
f.cum <- function(x, theta){pgamma(x, shape=theta[1], rate=theta[2])}
f.mode <- function(theta){shape <- theta[1]; rate <- theta[2]; mode <- ifelse(shape>1,(shape-1)/rate,NA); mode}
theta.from.moments <- function(m, v){shape <- m*m/v; rate <- m/v; c(shape, rate)}
dens.label <- 'dgamma'
parms.names <- c('shape', 'rate')
if (length(p) != 2) stop("Vector of probabilities p must be of length 2.")
if (length(q) != 2) stop("Vector of quantiles q must be of length 2.")
p <- sort(p); q <- sort(q)
#_____________________________________________________________________________________________________
print.area.text <- function(p, p.check, q, f, f.cum, F.inv, theta, mode, cex, plot.xlim, M=30, M0=50)
{
par.usr <- par('usr')
par.din <- par('din')
p.string <- as.character(round(c(0,1) + c(1,-1)*p.check, digits=4))
str.width <- strwidth(p.string, cex=cex)
str.height <- strheight("0", cex=cex)
J <- matrix(1, nrow=M0, ncol=1)
x.units.1in <- diff(par.usr[c(1,2)])/par.din[1]
y.units.1in <- diff(par.usr[c(3,4)])/par.din[2]
aspect.ratio <- y.units.1in/x.units.1in
# --- left area -----------------------------------------------------------
scatter.xlim <- c(max(plot.xlim[1], par.usr[1]), q[1])
scatter.ylim <- c(0, par.usr[4])
x <- seq(from=scatter.xlim[1], to=scatter.xlim[2], length=M)
y <- seq(from=scatter.ylim[1], to=scatter.ylim[2], length=M)
x.grid.index <- rep(seq(M), M)
y.grid.index <- rep(seq(M), rep(M, M))
grid.df <- f(x, theta)
# Estimate mass center
tmp.p <- seq(from=0, to=p[1], length=M0)
tmp.x <- F.inv(tmp.p, theta)
h <- f(tmp.x, theta)
mass.center <- c(mean(tmp.x), sum(h[-1]*diff(tmp.x))/diff(range(tmp.x)))
# Identify points under the curve
# (to eliminate them from the list of candidates)
gridpoint.under.the.curve <- y[y.grid.index] <= grid.df[x.grid.index]
w <- which(gridpoint.under.the.curve)
x <- x[x.grid.index]; y <- y[y.grid.index]
if (length(w)){x <- x[-w]; y <- y[-w]}
# Eliminate points to the right of the mode, if any
w <- which(x>mode)
if (length(w)){x <- x[-w]; y <- y[-w]}
# Eliminate points for which the text would fall out of the plot area
w <- which((par.usr[1]+str.width[1]) <= x & (y + str.height) <= par.usr[4])
x <- x[w]; y <- y[w]
# For each height, eliminate the closest point to the curve
# (we want to stay away from the curve to preserve readability)
w <- which(!duplicated(y, fromLast=T))
if (length(w)){x <- x[-w]; y <- y[-w]}
# For each point, compute distance from mass center and pick the closest point
d <- ((x-mass.center[1])^2) + ((y-mass.center[2])/aspect.ratio)^2
w <- which.min(d)
x <- x[w]; y <- y[w]
if (length(x))
{
text(x, y, labels=p.string[1], adj=c(1,0), col='gray', cex=cex)
}
else
{
text(plot.xlim[1], mean(par.usr[c(3,4)]), labels=p.string[1], col='gray', cex=cex, srt=90, adj=c(1,0))
}
# --- right area ----------------------------------------------------------
scatter.xlim <- c(q[2], plot.xlim[2])
scatter.ylim <- c(0, par.usr[4])
x <- seq(from=scatter.xlim[1], to=scatter.xlim[2], length=M)
y <- seq(from=scatter.ylim[1], to=scatter.ylim[2], length=M)
x.grid.index <- rep(seq(M), M)
y.grid.index <- rep(seq(M), rep(M, M))
grid.df <- f(x, theta)
# Estimate mass center
tmp.p <- seq(from=p[2], to=f.cum(plot.xlim[2], theta), length=M0)
tmp.x <- F.inv(tmp.p, theta)
h <- f(tmp.x, theta)
mass.center <- c(mean(tmp.x), sum(h[-length(h)]*diff(tmp.x))/diff(range(tmp.x)))
# Identify points under the curve
# (to eliminate them from the list of candidates)
gridpoint.under.the.curve <- y[y.grid.index] <= grid.df[x.grid.index]
w <- which(gridpoint.under.the.curve)
x <- x[x.grid.index]; y <- y[y.grid.index]
if (length(w)){x <- x[-w]; y <- y[-w]}
# Eliminate points to the left of the mode, if any
w <- which(x
if (length(w)){x <- x[-w]; y <- y[-w]}
# Eliminate points for which the text would fall out of the plot area
w <- which((par.usr[2]-str.width[2]) >= x & (y + str.height) <= par.usr[4])
x <- x[w]; y <- y[w]
# For each height, eliminate the closest point to the curve
# (we want to stay away from the curve to preserve readability)
w <- which(!duplicated(y))
if (length(w)){x <- x[-w]; y <- y[-w]}
# For each point, compute distance from mass center and pick the closest point
d <- ((x-mass.center[1])^2) + ((y-mass.center[2])/aspect.ratio)^2
w <- which.min(d)
x <- x[w]; y <- y[w]
if (length(x))
{
text(x, y, labels=p.string[2], adj=c(0,0), col='gray', cex=cex)
}
else
{
text(plot.xlim[2], mean(par.usr[c(3,4)]), labels=p.string[2], col='gray', cex=cex, srt=-90, adj=c(1,0))
}
}
# ......................................................................................................................................
Newton.Raphson <- function(derivative.epsilon, precision, f.cum, p, q, theta.from.moments, start.with.normal.approx, start)
{
Hessian <- matrix(NA, 2, 2)
if (start.with.normal.approx)
{
# Probably not a very good universal choice, but proved good in most cases in practice
m <- diff(q)/diff(p)*(0.5-p[1]) + q[1]
v <- (diff(q)/diff(qnorm(p)))^2
theta <- theta.from.moments(m, v)
}
else theta <- start
change <- precision + 1
niter <- 0
# Newton-Raphson multivariate algorithm
while (max(abs(change)) > precision)
{
Hessian[,1] <- (f.cum(q, theta) - f.cum(q, theta - c(derivative.epsilon, 0))) / derivative.epsilon
Hessian[,2] <- (f.cum(q, theta) - f.cum(q, theta - c(0, derivative.epsilon))) / derivative.epsilon
f <- f.cum(q, theta) - p
change <- solve(Hessian) %*% f
last.theta <- theta
theta <- last.theta - change
# If we step out of limits, reduce change
if (any(theta<0))
{
k <- min(last.theta/change)
theta <- last.theta - k/2*change
}
niter <- niter + 1
}
list(theta=as.vector(theta), niter=niter, last.change=as.vector(change))
}
# ...............................................................................................................
plot.density <- function(p, q, f, f.cum, F.inv, mode, theta, plot.xlim, dens.label, parms.names, cex)
{
if (length(plot.xlim) == 0)
{
plot.xlim <- F.inv(c(0, 1), theta)
if (is.infinite(plot.xlim[1]))
{
tmp <- min(c(0.001, p[1]/10))
plot.xlim[1] <- F.inv(tmp, theta)
}
if (is.infinite(plot.xlim[2]))
{
tmp <- max(c(0.999, 1 - (1-p[2])/10))
plot.xlim[2] <- F.inv(tmp, theta)
}
}
plot.xlim <- sort(plot.xlim)
x <- seq(from=min(plot.xlim), to=max(plot.xlim), length=1000)
h <- f(x, theta)
x0 <- x; f0 <- h
ylab <- paste(c(dens.label, '(x, ', parms.names[1], ' = ', round(theta[1], digits=5), ', ', parms.names[2], ' = ', round(theta[2], digits=5), ')'), collapse='')
plot(x, h, type='l', ylab=ylab)
# fill in area on the left side of the distribution
x <- seq(from=plot.xlim[1], to=q[1], length=1000)
y <- f(x, theta)
x <- c(x, q[1], plot.xlim[1]); y <- c(y, 0, 0)
polygon(x, y, col='lightgrey', border='lightgray')
# fill in area on the right side of the distribution
x <- seq(from=max(plot.xlim), to=q[2], length=1000)
y <- f(x, theta)
x <- c(x, q[2], plot.xlim[2]); y <- c(y, 0, 0)
polygon(x, y, col='lightgrey', border='lightgray')
# draw distrn again
points(x0, f0, type='l')
h <- f(q, theta)
points(rep(q[1], 2), c(0, h[1]), type='l', col='orange')
points(rep(q[2], 2), c(0, h[2]), type='l', col='orange')
# place text on both ends areas
print.area.text(p, p.check, q, f, f.cum, F.inv, theta, mode, cex, plot.xlim)
xaxp <- par("xaxp")
x.ticks <- seq(from=xaxp[1], to=xaxp[2], length=xaxp[3]+1)
q2print <- as.double(setdiff(as.character(q), as.character(x.ticks)))
mtext(q2print, side=1, col='orange', at=q2print, cex=0.6, line=2.1)
points(q, rep(par('usr')[3]+0.15*par('cxy')[2], 2), pch=17, col='orange')
}
#________________________________________________________________________________________________________________
parms <- Newton.Raphson(derivative.epsilon, precision, f.cum, p, q, theta.from.moments, start.with.normal.approx, start=start)
p.check <- f.cum(q, parms$theta)
if (plot) plot.density(p, q, f, f.cum, F.inv, f.mode(parms$theta), parms$theta, plot.xlim, dens.label, parms.names, 0.8)
list(shape=parms$theta[1], rate=parms$theta[2], scale=1/parms$theta[2], last.change=parms$last.change, niter=parms$niter, q=q, p=p, p.check=p.check)
}
precision=0.001, derivative.epsilon=1e-3, start.with.normal.approx=T, start=c(1.1, 1.1), plot=F, plot.xlim=numeric(0))
{
# Version 1.0.2 (December 2012)
#
# Function developed by
# Lawrence Joseph and Patrick B�lisle
# Division of Clinical Epidemiology
# Montreal General Hospital
# Montreal, Qc, Can
#
# patrick.belisle@rimuhc.ca
# http://www.medicine.mcgill.ca/epidemiology/Joseph/PBelisle/GammaParmsFromQuantiles.html
#
# Please refer to our webpage for details on each argument.
f <- function(x, theta){dgamma(x, shape=theta[1], rate=theta[2])}
F.inv <- function(x, theta){qgamma(x, shape=theta[1], rate=theta[2])}
f.cum <- function(x, theta){pgamma(x, shape=theta[1], rate=theta[2])}
f.mode <- function(theta){shape <- theta[1]; rate <- theta[2]; mode <- ifelse(shape>1,(shape-1)/rate,NA); mode}
theta.from.moments <- function(m, v){shape <- m*m/v; rate <- m/v; c(shape, rate)}
dens.label <- 'dgamma'
parms.names <- c('shape', 'rate')
if (length(p) != 2) stop("Vector of probabilities p must be of length 2.")
if (length(q) != 2) stop("Vector of quantiles q must be of length 2.")
p <- sort(p); q <- sort(q)
#_____________________________________________________________________________________________________
print.area.text <- function(p, p.check, q, f, f.cum, F.inv, theta, mode, cex, plot.xlim, M=30, M0=50)
{
par.usr <- par('usr')
par.din <- par('din')
p.string <- as.character(round(c(0,1) + c(1,-1)*p.check, digits=4))
str.width <- strwidth(p.string, cex=cex)
str.height <- strheight("0", cex=cex)
J <- matrix(1, nrow=M0, ncol=1)
x.units.1in <- diff(par.usr[c(1,2)])/par.din[1]
y.units.1in <- diff(par.usr[c(3,4)])/par.din[2]
aspect.ratio <- y.units.1in/x.units.1in
# --- left area -----------------------------------------------------------
scatter.xlim <- c(max(plot.xlim[1], par.usr[1]), q[1])
scatter.ylim <- c(0, par.usr[4])
x <- seq(from=scatter.xlim[1], to=scatter.xlim[2], length=M)
y <- seq(from=scatter.ylim[1], to=scatter.ylim[2], length=M)
x.grid.index <- rep(seq(M), M)
y.grid.index <- rep(seq(M), rep(M, M))
grid.df <- f(x, theta)
# Estimate mass center
tmp.p <- seq(from=0, to=p[1], length=M0)
tmp.x <- F.inv(tmp.p, theta)
h <- f(tmp.x, theta)
mass.center <- c(mean(tmp.x), sum(h[-1]*diff(tmp.x))/diff(range(tmp.x)))
# Identify points under the curve
# (to eliminate them from the list of candidates)
gridpoint.under.the.curve <- y[y.grid.index] <= grid.df[x.grid.index]
w <- which(gridpoint.under.the.curve)
x <- x[x.grid.index]; y <- y[y.grid.index]
if (length(w)){x <- x[-w]; y <- y[-w]}
# Eliminate points to the right of the mode, if any
w <- which(x>mode)
if (length(w)){x <- x[-w]; y <- y[-w]}
# Eliminate points for which the text would fall out of the plot area
w <- which((par.usr[1]+str.width[1]) <= x & (y + str.height) <= par.usr[4])
x <- x[w]; y <- y[w]
# For each height, eliminate the closest point to the curve
# (we want to stay away from the curve to preserve readability)
w <- which(!duplicated(y, fromLast=T))
if (length(w)){x <- x[-w]; y <- y[-w]}
# For each point, compute distance from mass center and pick the closest point
d <- ((x-mass.center[1])^2) + ((y-mass.center[2])/aspect.ratio)^2
w <- which.min(d)
x <- x[w]; y <- y[w]
if (length(x))
{
text(x, y, labels=p.string[1], adj=c(1,0), col='gray', cex=cex)
}
else
{
text(plot.xlim[1], mean(par.usr[c(3,4)]), labels=p.string[1], col='gray', cex=cex, srt=90, adj=c(1,0))
}
# --- right area ----------------------------------------------------------
scatter.xlim <- c(q[2], plot.xlim[2])
scatter.ylim <- c(0, par.usr[4])
x <- seq(from=scatter.xlim[1], to=scatter.xlim[2], length=M)
y <- seq(from=scatter.ylim[1], to=scatter.ylim[2], length=M)
x.grid.index <- rep(seq(M), M)
y.grid.index <- rep(seq(M), rep(M, M))
grid.df <- f(x, theta)
# Estimate mass center
tmp.p <- seq(from=p[2], to=f.cum(plot.xlim[2], theta), length=M0)
tmp.x <- F.inv(tmp.p, theta)
h <- f(tmp.x, theta)
mass.center <- c(mean(tmp.x), sum(h[-length(h)]*diff(tmp.x))/diff(range(tmp.x)))
# Identify points under the curve
# (to eliminate them from the list of candidates)
gridpoint.under.the.curve <- y[y.grid.index] <= grid.df[x.grid.index]
w <- which(gridpoint.under.the.curve)
x <- x[x.grid.index]; y <- y[y.grid.index]
if (length(w)){x <- x[-w]; y <- y[-w]}
# Eliminate points to the left of the mode, if any
w <- which(x
# Eliminate points for which the text would fall out of the plot area
w <- which((par.usr[2]-str.width[2]) >= x & (y + str.height) <= par.usr[4])
x <- x[w]; y <- y[w]
# For each height, eliminate the closest point to the curve
# (we want to stay away from the curve to preserve readability)
w <- which(!duplicated(y))
if (length(w)){x <- x[-w]; y <- y[-w]}
# For each point, compute distance from mass center and pick the closest point
d <- ((x-mass.center[1])^2) + ((y-mass.center[2])/aspect.ratio)^2
w <- which.min(d)
x <- x[w]; y <- y[w]
if (length(x))
{
text(x, y, labels=p.string[2], adj=c(0,0), col='gray', cex=cex)
}
else
{
text(plot.xlim[2], mean(par.usr[c(3,4)]), labels=p.string[2], col='gray', cex=cex, srt=-90, adj=c(1,0))
}
}
# ......................................................................................................................................
Newton.Raphson <- function(derivative.epsilon, precision, f.cum, p, q, theta.from.moments, start.with.normal.approx, start)
{
Hessian <- matrix(NA, 2, 2)
if (start.with.normal.approx)
{
# Probably not a very good universal choice, but proved good in most cases in practice
m <- diff(q)/diff(p)*(0.5-p[1]) + q[1]
v <- (diff(q)/diff(qnorm(p)))^2
theta <- theta.from.moments(m, v)
}
else theta <- start
change <- precision + 1
niter <- 0
# Newton-Raphson multivariate algorithm
while (max(abs(change)) > precision)
{
Hessian[,1] <- (f.cum(q, theta) - f.cum(q, theta - c(derivative.epsilon, 0))) / derivative.epsilon
Hessian[,2] <- (f.cum(q, theta) - f.cum(q, theta - c(0, derivative.epsilon))) / derivative.epsilon
f <- f.cum(q, theta) - p
change <- solve(Hessian) %*% f
last.theta <- theta
theta <- last.theta - change
# If we step out of limits, reduce change
if (any(theta<0))
{
k <- min(last.theta/change)
theta <- last.theta - k/2*change
}
niter <- niter + 1
}
list(theta=as.vector(theta), niter=niter, last.change=as.vector(change))
}
# ...............................................................................................................
plot.density <- function(p, q, f, f.cum, F.inv, mode, theta, plot.xlim, dens.label, parms.names, cex)
{
if (length(plot.xlim) == 0)
{
plot.xlim <- F.inv(c(0, 1), theta)
if (is.infinite(plot.xlim[1]))
{
tmp <- min(c(0.001, p[1]/10))
plot.xlim[1] <- F.inv(tmp, theta)
}
if (is.infinite(plot.xlim[2]))
{
tmp <- max(c(0.999, 1 - (1-p[2])/10))
plot.xlim[2] <- F.inv(tmp, theta)
}
}
plot.xlim <- sort(plot.xlim)
x <- seq(from=min(plot.xlim), to=max(plot.xlim), length=1000)
h <- f(x, theta)
x0 <- x; f0 <- h
ylab <- paste(c(dens.label, '(x, ', parms.names[1], ' = ', round(theta[1], digits=5), ', ', parms.names[2], ' = ', round(theta[2], digits=5), ')'), collapse='')
plot(x, h, type='l', ylab=ylab)
# fill in area on the left side of the distribution
x <- seq(from=plot.xlim[1], to=q[1], length=1000)
y <- f(x, theta)
x <- c(x, q[1], plot.xlim[1]); y <- c(y, 0, 0)
polygon(x, y, col='lightgrey', border='lightgray')
# fill in area on the right side of the distribution
x <- seq(from=max(plot.xlim), to=q[2], length=1000)
y <- f(x, theta)
x <- c(x, q[2], plot.xlim[2]); y <- c(y, 0, 0)
polygon(x, y, col='lightgrey', border='lightgray')
# draw distrn again
points(x0, f0, type='l')
h <- f(q, theta)
points(rep(q[1], 2), c(0, h[1]), type='l', col='orange')
points(rep(q[2], 2), c(0, h[2]), type='l', col='orange')
# place text on both ends areas
print.area.text(p, p.check, q, f, f.cum, F.inv, theta, mode, cex, plot.xlim)
xaxp <- par("xaxp")
x.ticks <- seq(from=xaxp[1], to=xaxp[2], length=xaxp[3]+1)
q2print <- as.double(setdiff(as.character(q), as.character(x.ticks)))
mtext(q2print, side=1, col='orange', at=q2print, cex=0.6, line=2.1)
points(q, rep(par('usr')[3]+0.15*par('cxy')[2], 2), pch=17, col='orange')
}
#________________________________________________________________________________________________________________
parms <- Newton.Raphson(derivative.epsilon, precision, f.cum, p, q, theta.from.moments, start.with.normal.approx, start=start)
p.check <- f.cum(q, parms$theta)
if (plot) plot.density(p, q, f, f.cum, F.inv, f.mode(parms$theta), parms$theta, plot.xlim, dens.label, parms.names, 0.8)
list(shape=parms$theta[1], rate=parms$theta[2], scale=1/parms$theta[2], last.change=parms$last.change, niter=parms$niter, q=q, p=p, p.check=p.check)
}
Top
Download
gamma.parms.from.quantiles is a free R function. Download version 1.0.2 now.
Top
See also
See also beta.parms.from.quantiles, an R function developped to compute Beta distribution parameters fitting specified quantiles and cumulative probabilities.